Ecology and Evolution of Infectious Diseases
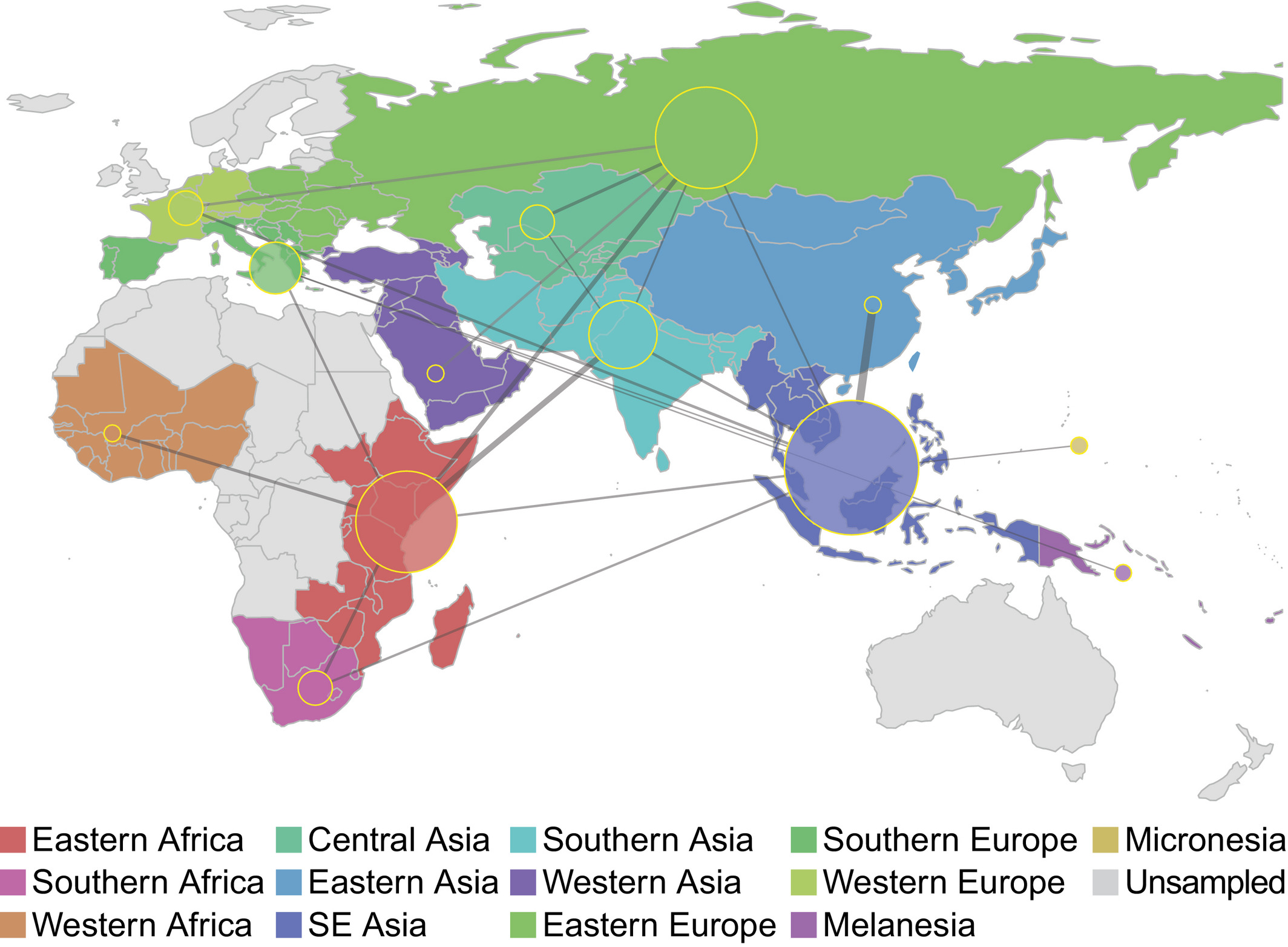
O'Neill, MB, Shockey, A, Zarley, A, et al. Lineage specific histories of Mycobacterium tuberculosis dispersal in Africa and Eurasia. Mol Ecol. 2019; 28: 3241– 3256.
Humans encounter billions of micro-organisms, and the vast majority of these interactions are inconsequential or beneficial to us. We are interested in how and why bacterial infections turn into diseases: What is the ecological function of virulence? How are genomes reshaped as bacteria shift from free-living to host associated lifestyles? Which adaptations enable host invasion? We use a range of approaches to address these questions, including analyses of population genomic and other data, in silico models, experimental evolution, and molecular genetics. Research in our lab focuses on several species of bacteria that represent the diversity of adaptive paths to virulence.